Lab 8: Spacing and distances
This session is concerned with summary statistics for spacings and interpoint distances. The lecturer’s R script is available here (right click and save).
library(spatstat)
Exercise 1
For the swedishpines data:
-
Calculate the estimate of the nearest neighbour distance distribution function G using
Gest.G <- Gest(swedishpines) -
Plot the estimate of G(r) against r
plot(G, cbind(km, rs, han) ~ r, main = "Nearest neighbor distance distribution")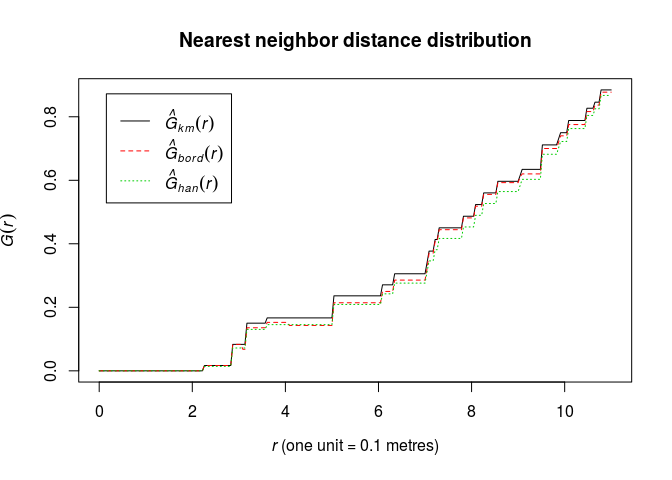
-
Plot the estimate of G(r) against the theoretical (Poisson) value Gpois(r)=1 − exp(−λπr2).
E.g.
plot(G, . ~ theo, main = "Nearest neighbor distribution")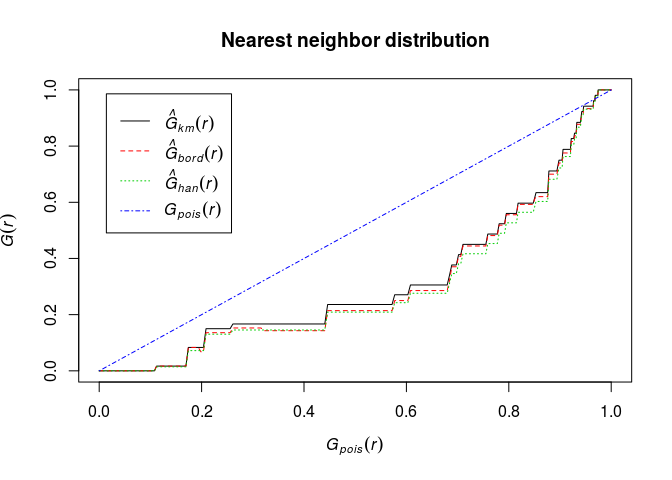
-
Define Fisher’s variance-stabilising transformation for c.d.f.’s by
Phi <- function(x) asin(sqrt(x))Plot the G function using the formula
Phi(.) ~ Phi(theo)and interpret it.Phi <- function(x) asin(sqrt(x)) plot(G, Phi(.) ~ Phi(theo), main = "Nearest neighbor distribution")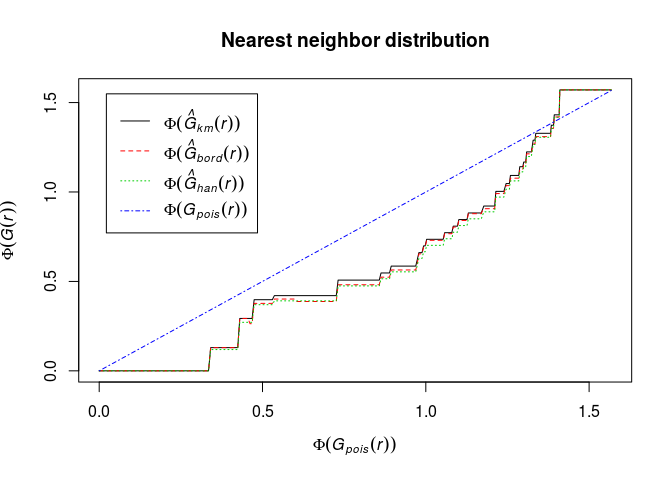
The transformation has made the deviations from CSR in the central part of the curve smaller. We need envelopes to say anything about significance.
Exercise 2
For the swedishpines data:
-
Calculate the estimate of the nearest neighbour distance distribution function F using
Fest.Fhat <- Fest(swedishpines) -
Plot the estimate of F(r) against r
plot(Fhat, main = "Empty Space function")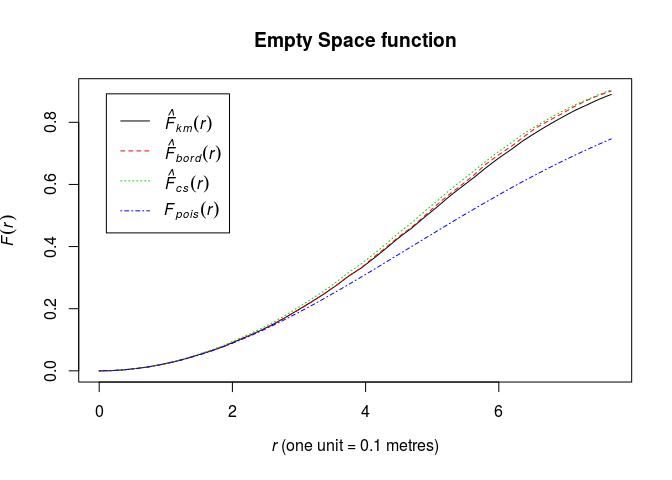
-
Plot the estimate of F(r) against the theoretical (Poisson) value Fpois(r)=1 − exp(−λπr2).
plot(Fhat, . ~ theo, main = "")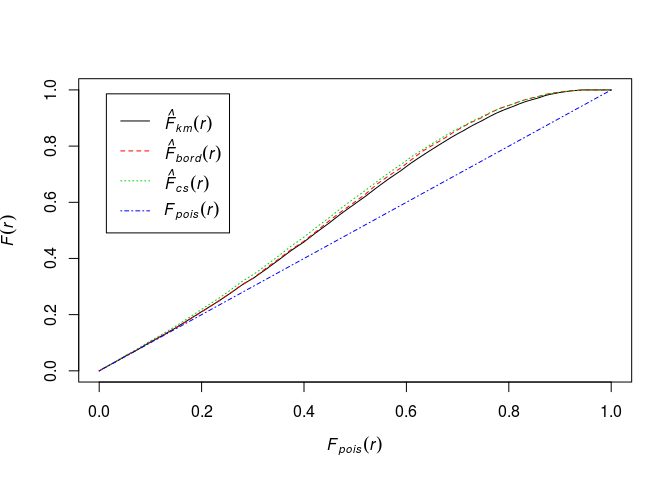
-
Define Fisher’s variance-stabilising transformation for c.d.f.’s by
Phi <- function(x) asin(sqrt(x))Plot the F function using the formula
Phi(.) ~ Phi(theo)and interpret it.Phi <- function(x) asin(sqrt(x)) plot(Fhat, Phi(.) ~ Phi(theo), main = "")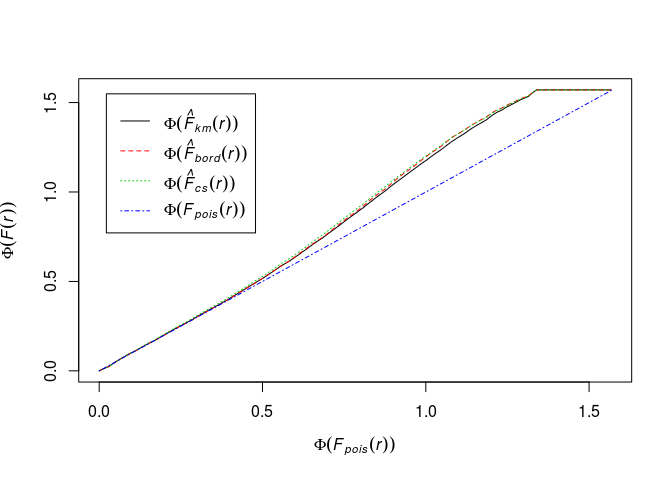
The transformation has changed the picture much. It looks like deviation from CSR, but again, without envelopes it’s hard to draw conclusions.